This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Transcriptomics?
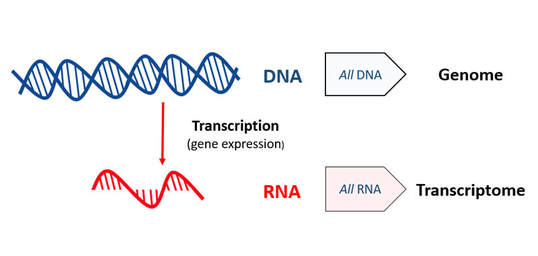
Transcriptomics is the study of the transcriptome—the complete set of RNA transcripts that are produced by the genome, under specific circumstances or in a specific cell—using high-throughput methods [1]. The two main methods are microarray analysis and RNA sequencing. Transcriptomics is specifically focused on how transcript patterns are affected by development, disease, or environmental factors such as hormones, drugs, etc [2]. Comparison of transcriptomes allows the identification of genes that are differentially expressed in distinct cell populations, or in response to different treatments [1].
What is a Microarray?
|
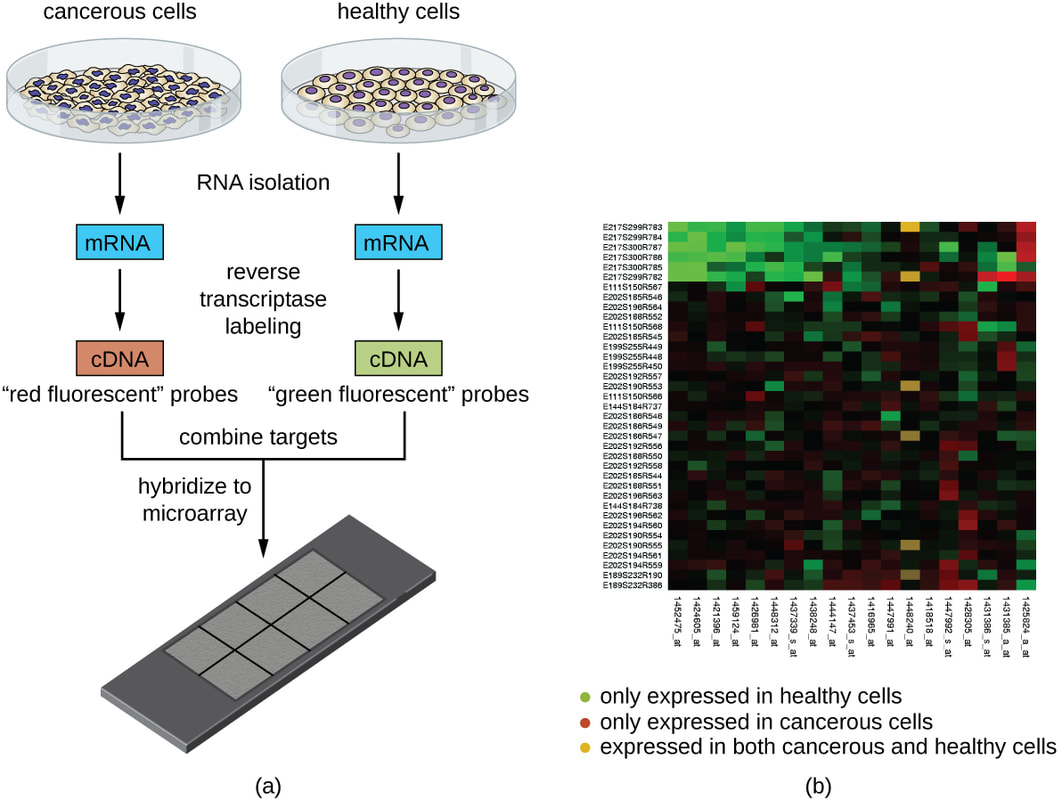
A microarray is a laboratory tool used to detect the expression of thousands of genes at the same time. DNA microarrays are microscope slides that are printed with thousands of tiny spots in defined positions, with each spot containing a known DNA sequence or gene. Often, these slides are referred to as gene chips or DNA chips. The DNA molecules attached to each slide act as probes to detect gene expression, which is also known as the transcriptome or the set of messenger RNA (mRNA) transcripts expressed by a group of genes. [3] |
What is Microarray Analysis?
What is a RNA Sequencing?
|
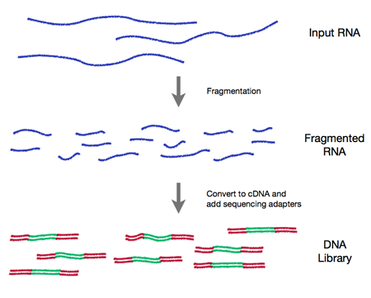
RNA sequencing is rapidly replacing gene expression microarrays in many labs. RNA-seq lets you quantify, discover and profile RNAs. For this technique, mRNA (and other RNAs) are first converted to cDNA. The cDNA is then used as the input for a next-generation sequencing library preparation. [5] |
The Human Protein Atlas
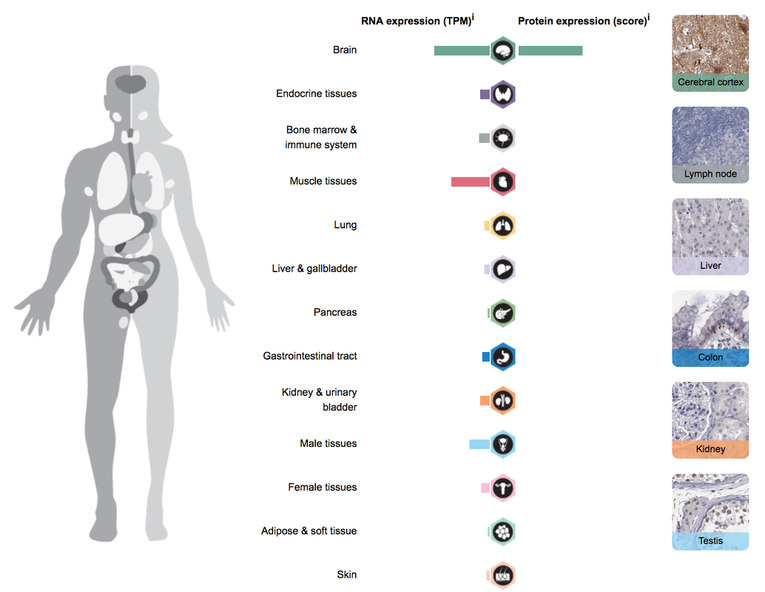
The Human Protein Atlas (HPA) is a program with the aim to map all the human proteins in cells, tissues and organs using integration of various omics technologies, including antibody-based imaging, mass spectrometry-based proteomics, transcriptomics and systems biology [6]. The Human Protein Atlas consists of three separate parts, each focusing on a particular aspect of the genome-wide analysis of the human proteins; the Tissue Atlas showing the distribution of the proteins across all major tissues and organs in the human body, the Cell Atlas showing the subcellular localization of proteins in single cells, and finally the Pathology Atlas showing the impact of protein levels for survival of patients with cancer [6].
Conclusion
Transcriptomics is useful in mapping out human proteins in cells. By using the Human Protein Atlas (HPA), I was able to discover in what tissues the NRXN3 protein is most prevalent in. From the results above, it is seen that RNA and Protein expression is very high in the brain, more specifically the cerebral cortex which contains neuronal cell bodies. The NRXN3 protein is not expressed anywhere else besides the brain, meaning it functions solely in the brain. However, RNA expression is also present in muscle tissue and interestingly enough male tissues. It appears that In men, but not women, having a certain variant of NRXN3 increased the risk of problems with alcohol by 2.5 times [7]. Seeing as though NRXN3 expression in women is significantly lower, this would be rather fascinating to follow up on with future research.
References
[1] Nature. (n.d.). Transcriptomics. Retrieved March 12, 2018, from https://www.nature.com/subjects/transcriptomics
[2] (n.d.). Retrieved from http://omicscentre.com/services-applications/genomics-and-transcriptomics/
[3] (n.d.). Retrieved from https://www.nature.com/scitable/definition/microarray-202
[4] DNA Microarray. (n.d.). Retrieved from http://learn.genetics.utah.edu/content/labs/microarray/
[5] An Introduction to RNA-seq. (2017, December 07). Retrieved from https://bitesizebio.com/13542/what-everyone-should-know-about-rna-seq/
[6] Introduction. (n.d.). Retrieved from https://www.proteinatlas.org/about
[7] Stoltenberg, S. F., Lehmann, M. K., Christ, C. C., Hersrud, S. L., & Davies, G. E. (2011). Associations among types of impulsivity, substance use problems and Neurexin-3 polymorphisms. Drug and Alcohol Dependence, 119(3). doi:10.1016/j.drugalcdep.2011.05.025
Images
Header: https://www.med24.ee/static/files/057/shutterstock_mikro-rna_mirna.jpg
https://deutsch.physics.ucsc.edu/microarray.gif
Figure 1: http://weclipart.com/gimg/AC1F2909E73C37C8/trascriptome.jpg
Figure 2: https://upload.wikimedia.org/wikipedia/commons/a/ae/OSC_Microbio_12_02_Microarray.jpg
Figure 3: https://rnaseq.uoregon.edu/img/fig-rna-seq.png
Figure 4: https://www.proteinatlas.org/ENSG00000021645-NRXN3/tissue
[1] Nature. (n.d.). Transcriptomics. Retrieved March 12, 2018, from https://www.nature.com/subjects/transcriptomics
[2] (n.d.). Retrieved from http://omicscentre.com/services-applications/genomics-and-transcriptomics/
[3] (n.d.). Retrieved from https://www.nature.com/scitable/definition/microarray-202
[4] DNA Microarray. (n.d.). Retrieved from http://learn.genetics.utah.edu/content/labs/microarray/
[5] An Introduction to RNA-seq. (2017, December 07). Retrieved from https://bitesizebio.com/13542/what-everyone-should-know-about-rna-seq/
[6] Introduction. (n.d.). Retrieved from https://www.proteinatlas.org/about
[7] Stoltenberg, S. F., Lehmann, M. K., Christ, C. C., Hersrud, S. L., & Davies, G. E. (2011). Associations among types of impulsivity, substance use problems and Neurexin-3 polymorphisms. Drug and Alcohol Dependence, 119(3). doi:10.1016/j.drugalcdep.2011.05.025
Images
Header: https://www.med24.ee/static/files/057/shutterstock_mikro-rna_mirna.jpg
https://deutsch.physics.ucsc.edu/microarray.gif
Figure 1: http://weclipart.com/gimg/AC1F2909E73C37C8/trascriptome.jpg
Figure 2: https://upload.wikimedia.org/wikipedia/commons/a/ae/OSC_Microbio_12_02_Microarray.jpg
Figure 3: https://rnaseq.uoregon.edu/img/fig-rna-seq.png
Figure 4: https://www.proteinatlas.org/ENSG00000021645-NRXN3/tissue