This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are Protein Domains?
Protein domains are distinct functional and/or structural units in a protein. They are usually responsible for a particular function or interaction, contributing to the overall role of a protein. Domains may exist in a variety of biological contexts, where similar domains can be found in proteins with different functions. [1]
Domains Found in NRXN3
Protein domains were found in NRXN3 using domain analysis tools such as Pfam, SMART, InterPRo, and PROSITE. All websites were utilized in order to make sure that conflicting results in domains found could be taken into consideration.
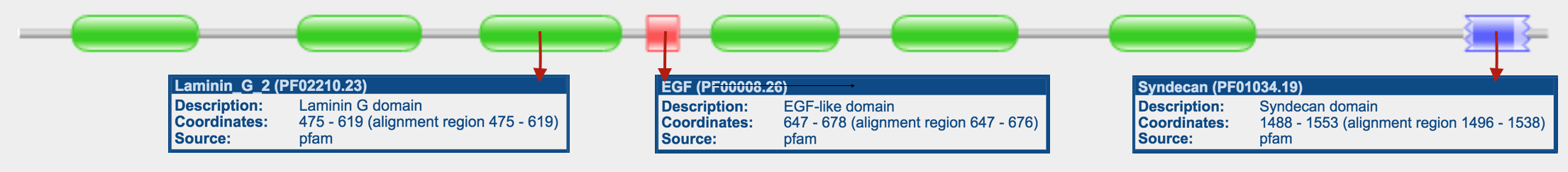
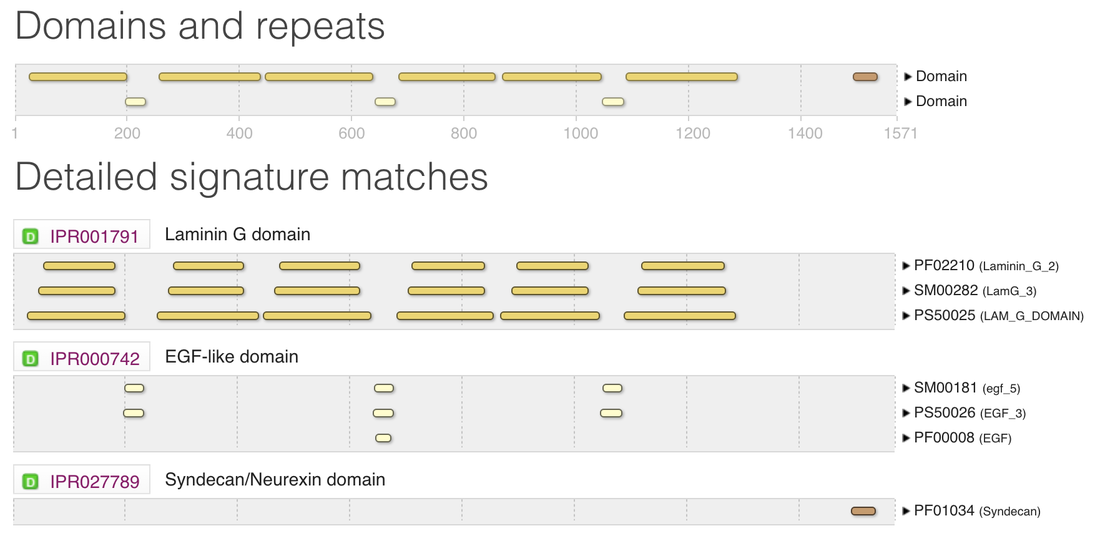
Pfam, SMART, InterPRo, and PROSITE were used in order to search for protein domains found in TCF4. The different sites were used in order to evaluate how consistently protein domains were identified across programs. The outputs for TCF4 were quite consistent across the platforms. Results from Pfam, SMART, and InterPRo are shown in the figures below:
Pfam, SMART, InterPRo, and PROSITE were used in order to search for protein domains found in TCF4. The different sites were used in order to evaluate how consistently protein domains were identified across programs. The outputs for TCF4 were quite consistent across the platforms. Results from Pfam, SMART, and InterPRo are shown in the figures below:
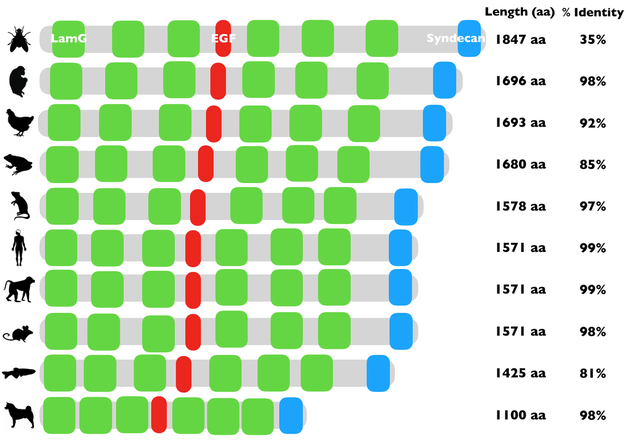
There is always a chance that results may differ from based on which tool you use to search with. That is why it is crucial that you take the time to investigate each site to see if you obtain similar or different results. Pfam, InterPRo, and SMART showed 3 main protein domains in NRXN3. Those domains are the Laminin G domain, EGF-like domain, and Syndecan domain.
|
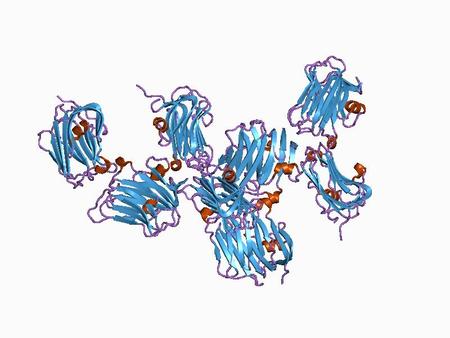
Laminin G (LamG) Domain The laminin globular (G) domain, also known as the LNS (Laminin-alpha, Neurexin and Sex hormone-binding globulin) domain, is on average 177 amino acids in length and can be found in one to six copies in various laminin family members as well as in a large number of other extracellular proteins. [2]
The exact function of the Laminin G domains has remained elusive, and a variety of binding functions has been ascribed to different Laminin G modules. Laminin G-containing proteins appear to have a wide variety of roles in cell adhesion, signaling, migration, assembly, and differentiation. [2] |
|
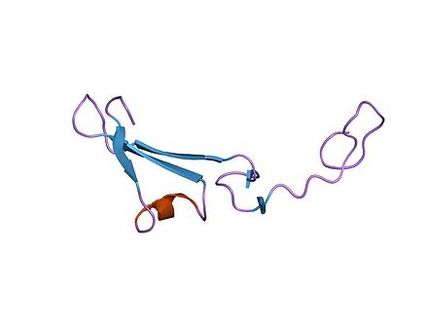
EGF-like Domain The EGF-like domain is an evolutionary conserved protein domain, which derives its name from the epidermal growth factor where it was first described. The EGF-like domain is not required for maximal cellular adhesion in L-selectin (expressed on lymphocytes). However, it is involved in both ligand recognition and adhesion in P-selectin (expressed on platelets) and may also be involved in protein-protein interactions. [3]
The EGF-like domain appears to play a vital role in immune responses and in eliminating dead cells in the organism. [3] |
|
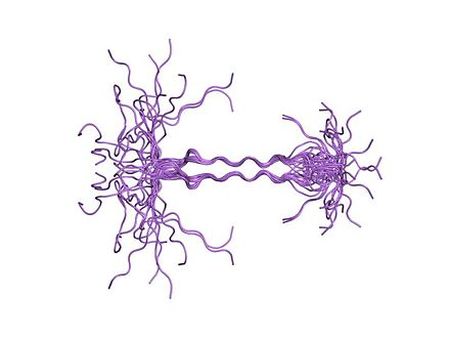
Syndecan Domain Syndecans are single transmembrane domain proteins that are thought to act as coreceptors, especially for G protein-coupled receptors. Syndecans are expressed on the cell surface in a cell-specific manner. The syndecan protein family has four members. Syndecans 1 and 3 and syndecans 2 and 4 make up separate subfamilies. Syndecan 3 is highly expressed in neural cells, but has low or undetectable amount in epithelial cells. In tissues, it is specific to the brain and expressed at low levels in liver, kidney, lung and small intestine. The functions of syndecan domain proteins include growth-factor-receptor activation, matrix adhesion, cell–cell adhesion, and tumor suppression and progression. [4]
|
Conservation of Domains in NRXN3
The Laminin G domain, EGF-like domain, and Syndecan domain are well conserved in the homologs that were selected for this study.
Conclusion
Three NRXN3 protein domains were found that were well conserved across homologous protein sequences in several organisms. Those three domains were the Laminin G domain, EFG-like domain, and Syndecan domain. The conservation of each domain among homologs shows that each domain plays a crucial role in the function and or structure of the protein. Cell-adhesion appears to be a function that is shared between all three domains making them key players in the role of the protein.
References
[1] Protein Classification. (2016, June 08). Retrieved March 15, 2018, from https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification
[2] http://pfam.xfam.org/family/PF02210.23
[3] http://pfam.xfam.org/family/PF00008.26
[4] http://pfam.xfam.org/family/PF01034.19
Images
Header: https://nl.wikipedia.org/wiki/Cytochroom-c-oxidase
Figure 1: http://pfam.xfam.org/search/sequence
Figure 2: http://smart.embl-heidelberg.de/
Figure 3: http://www.ebi.ac.uk/interpro/
Figure 4: https://en.wikipedia.org/wiki/File:PDB_1c4r_EBI.jpg
Figure 5: https://en.wikipedia.org/wiki/File:PDB_1hre_EBI.jpg
Figure 6: https://en.wikipedia.org/wiki/File:PDB_1ejq_EBI.jpg
[1] Protein Classification. (2016, June 08). Retrieved March 15, 2018, from https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification
[2] http://pfam.xfam.org/family/PF02210.23
[3] http://pfam.xfam.org/family/PF00008.26
[4] http://pfam.xfam.org/family/PF01034.19
Images
Header: https://nl.wikipedia.org/wiki/Cytochroom-c-oxidase
Figure 1: http://pfam.xfam.org/search/sequence
Figure 2: http://smart.embl-heidelberg.de/
Figure 3: http://www.ebi.ac.uk/interpro/
Figure 4: https://en.wikipedia.org/wiki/File:PDB_1c4r_EBI.jpg
Figure 5: https://en.wikipedia.org/wiki/File:PDB_1hre_EBI.jpg
Figure 6: https://en.wikipedia.org/wiki/File:PDB_1ejq_EBI.jpg