This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Alcohol Dependence and NRXN3
Alcohol dependence is a serious public health issue and is described as a strong, often uncontrollable, desire to drink [1]. Alcohol is the most commonly used addictive substance in the United States: 17.6 million people, or one in every 12 adults, suffer from alcohol abuse or dependence along with several million more who engage in risky, binge drinking patterns that could lead to alcohol problems [2]. Signs of alcohol dependence include experiencing blackouts and short-term memory loss, extreme mood swings, making excuses for drinking, drinking in inappropriate settings, lack of control, and withdrawal symptoms [3].
In addition to social and environmental factors, alcohol dependence is associated with a genetic contribution and one such gene, NRXN3 was recently identified. NRXN3 belongs to a family of neurexin presynaptic transmembrane proteins that function as cell adhesion molecules, connect neurons, and mediate cell signaling between neurons [4]. Polymorphisms of the NRXN3 gene distinguish individuals who are dependent on illegal substances from control individuals [5].
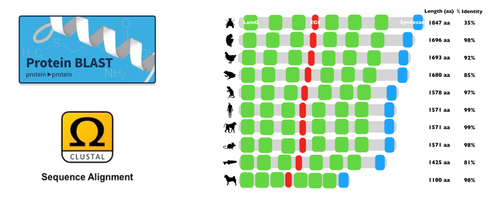
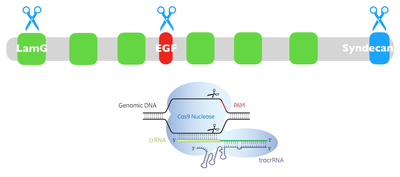
NRXN3 is generally well conserved among species and has three main protein domains. Those three domains are the Laminin G domain, EFG-like domain, and Syndecan domain. The conservation of each domain among homologs shows that each domain plays a crucial role in the function and or structure of the protein. Cell-adhesion appears to be a function that is shared between all three domains making them important in the role of the protein.
In addition to social and environmental factors, alcohol dependence is associated with a genetic contribution and one such gene, NRXN3 was recently identified. NRXN3 belongs to a family of neurexin presynaptic transmembrane proteins that function as cell adhesion molecules, connect neurons, and mediate cell signaling between neurons [4]. Polymorphisms of the NRXN3 gene distinguish individuals who are dependent on illegal substances from control individuals [5].
NRXN3 is generally well conserved among species and has three main protein domains. Those three domains are the Laminin G domain, EFG-like domain, and Syndecan domain. The conservation of each domain among homologs shows that each domain plays a crucial role in the function and or structure of the protein. Cell-adhesion appears to be a function that is shared between all three domains making them important in the role of the protein.
CASK and NRXN3
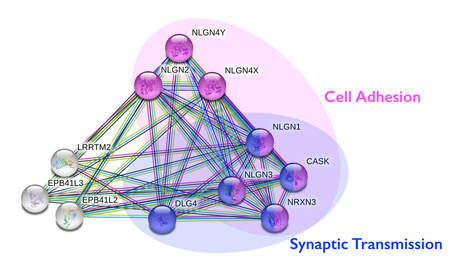
The protein interaction network above that was created with String was split into two main groups of proteins that shared similar functions. The group of pink proteins are all involved in cell adhesion. These proteins include neuroligins (NLGN4Y, NLGN2, NLGN4X, NLGN1, NLGN3), CASK (Calcium/calmodulin-dependent serine protein kinase), and NRXN3. It is expected that neuroligins and neurexins are involved in cell adhesion since they are cell adhesion molecules. The group of blue proteins are all involved in synaptic transmission. These proteins include neuroligins (NLGN1 and NLGN3), DLG4 (Discs Large MAGUK Scaffold Protein 4), CASK, and NRXN3. CASK and NRXN3 are both involved in synaptic transmission and cell adhesion. CASK is a serine protein kinase that helps control expression of other genes involved in brain development and phosphorylates neurexins.
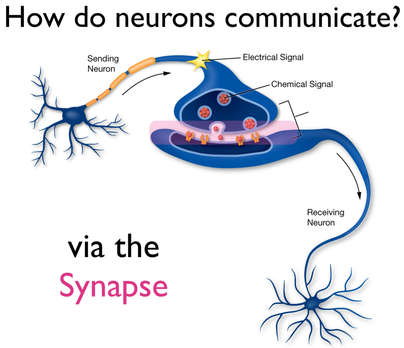
The Synapse is a Key PlayerNRXN3 is located specifically on the pre-synaptic membrane. The molecular function of NRXN3 is cell adhesion while an important biological process of the gene is synaptogenesis. Neurexins are cell adhesion molecules and when they function improperly, they aren't able to connect neurons together properly. This means that a synapse isn't able to form for the signals to pass through, resulting in the inability for information to pass from one neuron to the next. Therefore, synaptic function appears to play a crucial role in how NRXN3 protein dysfunction and alcohol dependence are related.
|
Goals and Hypothesis
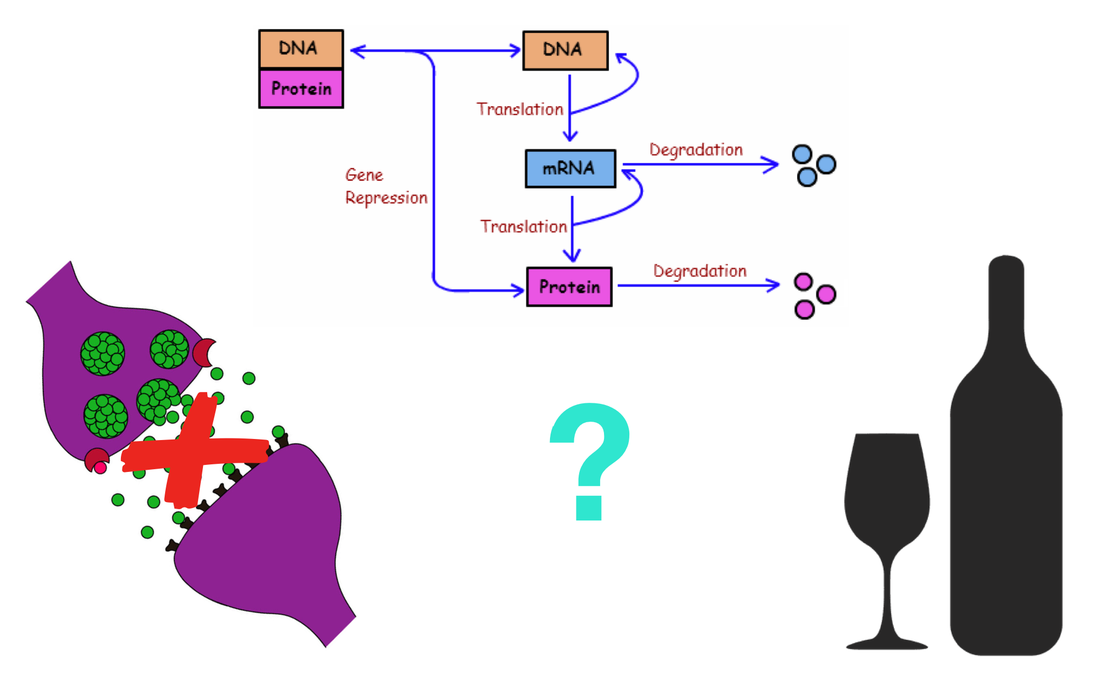
My primary goal is to better understand how NRXN3 mRNA regulation contributes to synaptic dysfunction and ultimately alcohol dependence. My long-term goal of this research is to understand how NRXN3 contributes to alcohol dependence so that effective treatments can be made for those who suffer from the disorder. My hypothesis is that low mRNA levels might be due to a decrease in phosphorylation at serine sites in NRXN3, leading to loss of protein function at the presynaptic transmembrane and synaptic dysfunction.
Model Organism
|
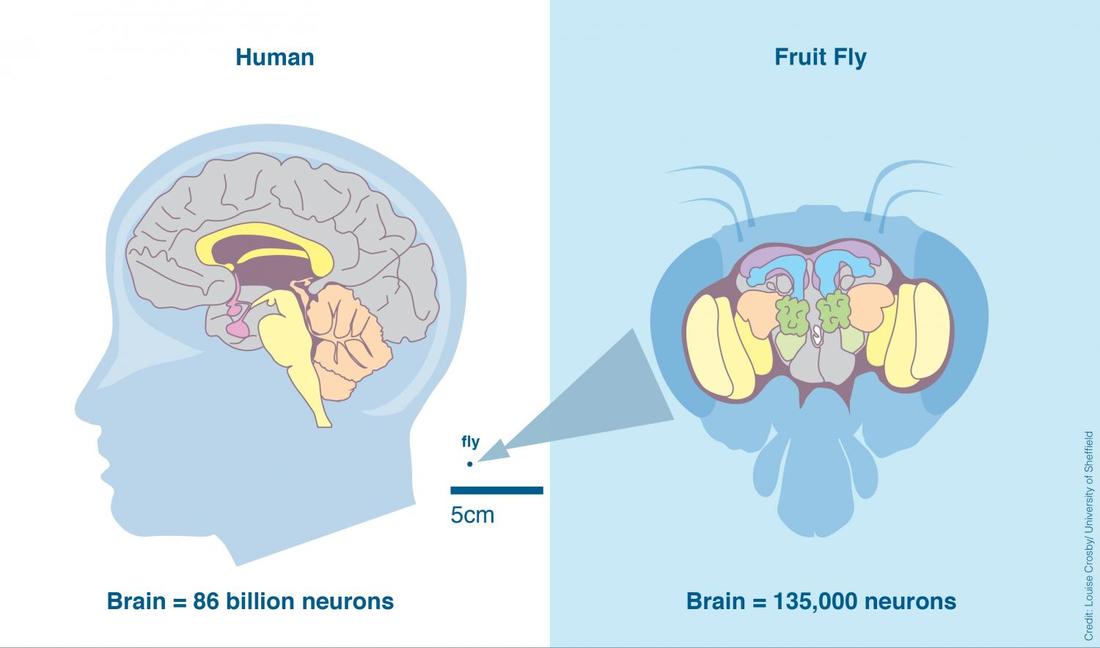
Drosophila melanogaster will be used as a model organism to study NRXN3 because they share anatomical and molecular similarities to humans. Drosophila’s synapse is useful to investigate due to its structural accessibility and because their nerve impulses direct locomotion and behavior like in humans. They are cheap to breed and easy to maintain in a lab setting. They have a short lifespan and fast maturation rate. Their genome is fully sequenced and are extensively used in brain research. They exhibit similar behavioral phenotypes to humans such as uncoordination when exposed to alcohol vapors. Therefore, it can be deduced that mutations in NRXN3 should induce the uncoordinated phenotype in Drosophila and indicate improper synaptic function and alcohol dependence.
|
Aim 1: Characterize conserved amino acids of NRXN3 crucial for proper synaptic function
I will use BLAST to gather FASTA animo acid sequences for homologs of NRXN3. I will then use Clustal Omega for Multiple Sequence Alignment and align those protein sequences in order to identify and determine well-conserved amino acid regions in NRXN3 among those homologs. I will then mutate those well-conserved amino acids with CRISPR-Cas9 using Drosophila as a model to understand how these mutations affect synaptic function and alcohol dependence at different stages of development. I will then screen the mutated flies that exhibit the incoordination phenotype as well as improper synaptic function and alcohol dependence.
My rationale behind this approach is that screening flies with the induced gene mutations from those conserved regions should result in uncoordinated flies. This in turn will then explain how mutations in NRXN3 exactly affect synaptic function.
I hypothesize that specific conserved amino acids in NRXN3 correlate with proper synaptic function and decreased alcohol dependence.
I hypothesize that specific conserved amino acids in NRXN3 correlate with proper synaptic function and decreased alcohol dependence.
Aim 2: Identify small molecules that rescue NRXN3 mutant phenotypes.
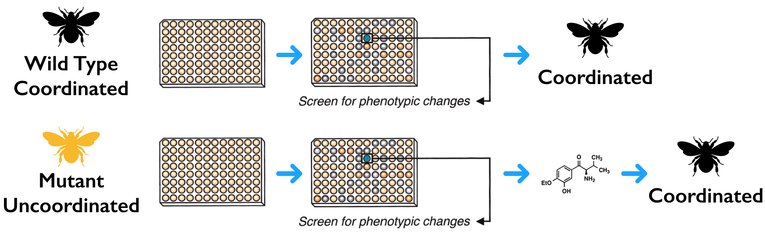
I will be performing a high-throughput chemical genomic screen on wild type flies and NRXN3 mutant flies from Aim 1. I will do this by using a diversity-oriented library to identify small molecules that will rescue the NRXN3 mutant incoordination and improper synaptic function phenotypes and restore proper synaptic function in the mutant fly.
My rationale behind this approach is that by treating mutant NRXN3 flies with the identified small molecules from the chemical genomic screen, cell adhesion will be up-regulated and the wild type function of NRXN3 proper synaptic function will be restored.
I hypothesize that small molecules that can rescue the NRXN3 mutant phenotypes will restore proper synaptic function and decrease alcohol dependence.
I hypothesize that small molecules that can rescue the NRXN3 mutant phenotypes will restore proper synaptic function and decrease alcohol dependence.
Aim 3: Identify and mutate phosphorylation sites in NRXN3 important for synaptic function
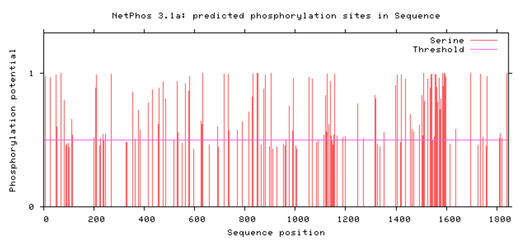
By using STRING, I will find proteins that interact with NRXN3 and are involved in phosphorylation. I will then use NetPhos to find the expected phosphorylation sites in NRXN3. I will specifically look for sites where serine is phosphorylated.
I will then mutate those serine phosphorylation sites in a wildtype NRXN3 fly and compare the phenotype of that mutant fly with a wildtype fly to observe how phosphorylation affects synaptic function.
My rationale behind this approach is that I previously found out that CASK is a serine protein kinase that helps control expression of other genes involved in brain development. It makes sense to look at potential serine sites because CASK phosphorylates serine sites in neurexins. Mutating the most highly conserved serine phosphorylation sites in NRXN3 should result in decreased levels of CASK activity, a decrease in phosphorylation at the serine sites, improper synaptic function, and alcohol dependence.
Future Directions
Future directions involve looking a bit further into the Syndecan domain because there appears to be a cluster of predicted serine sites around the same region as that domain. This leads me to believe that the Syndecan domain would be an interesting region to specifically research in the future. I would also look more into the the DLG4 protein that appears to interact with NRXN3 in the protein interaction network above and how mutating that protein would affect synaptic function as well. The end goal would be to identify compounds that can rescue the mutant phenotype and develop novel drugs to aid in preventing or treating alcohol dependent individuals.
References
[1] Alcohol dependence and withdrawal. (n.d.). Retrieved February 15, 2018, from https://www.drinkaware.co.uk/alcohol-facts/health-effects-of-alcohol/mental-health/alcohol-dependence/
[2] Wilcox, S. (n.d.). Facts About Alcohol. Retrieved from https://www.ncadd.org/about-addiction/alcohol/facts-about-alcohol
[3] Galbicsek, C. (n.d.). Signs of Alcoholism - Know the Warning Signs of Alcohol Abuse. Retrieved February 06, 2018, from https://www.alcoholrehabguide.org/alcohol/warning-signs/
[4] Stoltenberg, S. F., Lehmann, M. K., Christ, C. C., Hersrud, S. L., & Davies, G. E. (2011, December 15). Associations among types of impulsivity, substance use problems and Neurexin-3 polymorphisms. Retrieved February 15, 2018, from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3254149/
[5] Hishimoto, A., Liu, Q., Drgon, T., Pletnikova, O., Walther, D., Zhu, X., . . . Uhl, G. R. (2007, September 04). Neurexin 3 polymorphisms are associated with alcohol dependence and altered expression of specific isoforms | Human Molecular Genetics | Oxford Academic. Retrieved February 15, 2018, from https://academic.oup.com/hmg/article/16/23/2880/556881
Images
Header: https://ak0.picdn.net/shutterstock/videos/5171780/thumb/1.jpg
Figure 1: https://string-db.org/
Figure 2: http://learn.genetics.utah.edu/content/neuroscience/neurons/neurons-2.jpg
Figure 3: https://biology.tutorvista.com/cell/gene-regulation.html
Figure 3: https://images.vexels.com/media/users/3/137130/isolated/preview/7411ac190a0efd344066e2c01bc17c36-wine-bottle-glass-silhouette-by-vexels.png
Figure 3: https://static1.squarespace.com/static/52ec8c1ae4b047ccc14d6f29/t/5705bcb60442628269048e4b/1490436869202/synapse1.jpg
Figure 4: https://3c1703fe8d.site.internapcdn.net/newman/gfx/news/hires/2016/10-newresearchc.jpg
Figure 5: https://blast.ncbi.nlm.nih.gov/images/protein-blast-cover.png
Figure 5: http://www.clustal.org/images/clustalo_big.png
Figure 6: http://dharmacon.horizondiscovery.com/uploadedImages/Products/Gene_Editing/_assets/crtracr-diagram.jpg
Figure 7: http://www.plantphysiol.org/content/plantphysiol/133/2/448/F1.large.jpg?width=800&height=600&carousel=1
[1] Alcohol dependence and withdrawal. (n.d.). Retrieved February 15, 2018, from https://www.drinkaware.co.uk/alcohol-facts/health-effects-of-alcohol/mental-health/alcohol-dependence/
[2] Wilcox, S. (n.d.). Facts About Alcohol. Retrieved from https://www.ncadd.org/about-addiction/alcohol/facts-about-alcohol
[3] Galbicsek, C. (n.d.). Signs of Alcoholism - Know the Warning Signs of Alcohol Abuse. Retrieved February 06, 2018, from https://www.alcoholrehabguide.org/alcohol/warning-signs/
[4] Stoltenberg, S. F., Lehmann, M. K., Christ, C. C., Hersrud, S. L., & Davies, G. E. (2011, December 15). Associations among types of impulsivity, substance use problems and Neurexin-3 polymorphisms. Retrieved February 15, 2018, from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3254149/
[5] Hishimoto, A., Liu, Q., Drgon, T., Pletnikova, O., Walther, D., Zhu, X., . . . Uhl, G. R. (2007, September 04). Neurexin 3 polymorphisms are associated with alcohol dependence and altered expression of specific isoforms | Human Molecular Genetics | Oxford Academic. Retrieved February 15, 2018, from https://academic.oup.com/hmg/article/16/23/2880/556881
Images
Header: https://ak0.picdn.net/shutterstock/videos/5171780/thumb/1.jpg
Figure 1: https://string-db.org/
Figure 2: http://learn.genetics.utah.edu/content/neuroscience/neurons/neurons-2.jpg
Figure 3: https://biology.tutorvista.com/cell/gene-regulation.html
Figure 3: https://images.vexels.com/media/users/3/137130/isolated/preview/7411ac190a0efd344066e2c01bc17c36-wine-bottle-glass-silhouette-by-vexels.png
Figure 3: https://static1.squarespace.com/static/52ec8c1ae4b047ccc14d6f29/t/5705bcb60442628269048e4b/1490436869202/synapse1.jpg
Figure 4: https://3c1703fe8d.site.internapcdn.net/newman/gfx/news/hires/2016/10-newresearchc.jpg
Figure 5: https://blast.ncbi.nlm.nih.gov/images/protein-blast-cover.png
Figure 5: http://www.clustal.org/images/clustalo_big.png
Figure 6: http://dharmacon.horizondiscovery.com/uploadedImages/Products/Gene_Editing/_assets/crtracr-diagram.jpg
Figure 7: http://www.plantphysiol.org/content/plantphysiol/133/2/448/F1.large.jpg?width=800&height=600&carousel=1
Drafts
| kurt4-5-18finaltalk_draft1.pptx | |
| File Size: | 5444 kb |
| File Type: | pptx |
| kurt4-26-18finaltalk_finaldraft.pdf | |
| File Size: | 3115 kb |
| File Type: | |