Interaction Networks |
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein interaction networks?
Protein–protein interaction networks are the networks of protein complexes formed by biochemical events and/or electrostatic forces and that serve a distinct biological function as a complex. The protein interactome describes the full repertoire of a biological system’s protein–protein interactions (PPIs) [1]. The lines that connect proteins within these interaction networks show which proteins interact with each other. Databases such as String, IntAct, and BioGrid can be used to explore PPIs for a protein of interest and construct protein interaction networks for said protein.
NRXN3 Interaction Network
There are many ways in which you can generate a protein interaction network. Databases such as String, IntAct, and BioGrid can be utilized to explore the PPIs for your protein of interest. In my case, I used String to create a protein interaction network for NRXN3 because it had more functions and customization options than the other database tools did.
Conclusion
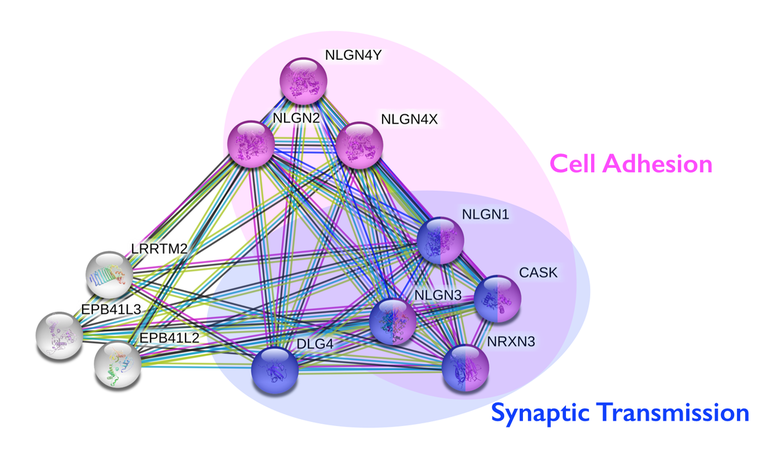
The protein interaction network above that was created with String was split into two main groups of proteins that shared similar functions. The group of pink proteins are all involved in cell adhesion. These proteins include neuroligins (NLGN4Y, NLGN2, NLGN4X, NLGN1, NLGN3), CASK (Calcium/calmodulin-dependent serine protein kinase), and NRXN3. It is expected that neuroligins and neurexins are involved in cell adhesion since they are cell adhesion molecules. The group of blue proteins are all involved in synaptic transmission. These proteins include neuroligins (NLGN1 and NLGN3), DLG4 (Discs Large MAGUK Scaffold Protein 4), CASK, and NRXN3. What is interesting is that CASK and NRXN3 are both involved in synaptic transmission and cell adhesion. CASK is a serine protein kinase that helps control expression of other genes involved in brain development. Since CASK phosphorylates Neurexins, mutating NRXN3 should result in decreased levels of CASK activity, a decrease in phosphorylation at the serine sites, and improper synaptic function.
References
[1] (n.d.). Retrieved April 13, 2018, from https://www.nature.com/subjects/protein-protein-interaction-networks
Images
Figure 1: https://string-db.org/
[1] (n.d.). Retrieved April 13, 2018, from https://www.nature.com/subjects/protein-protein-interaction-networks
Images
Figure 1: https://string-db.org/